Software
inStrain
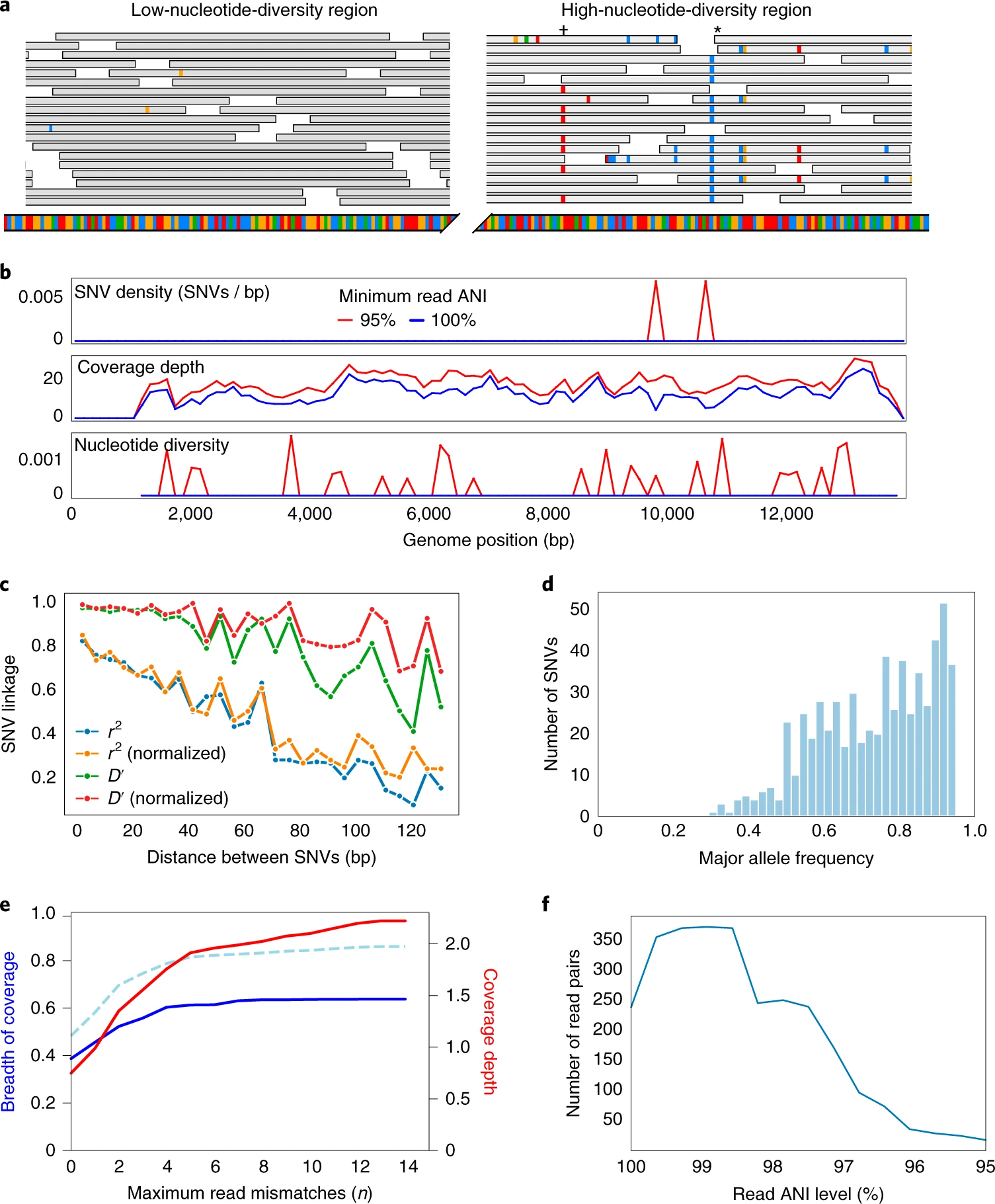
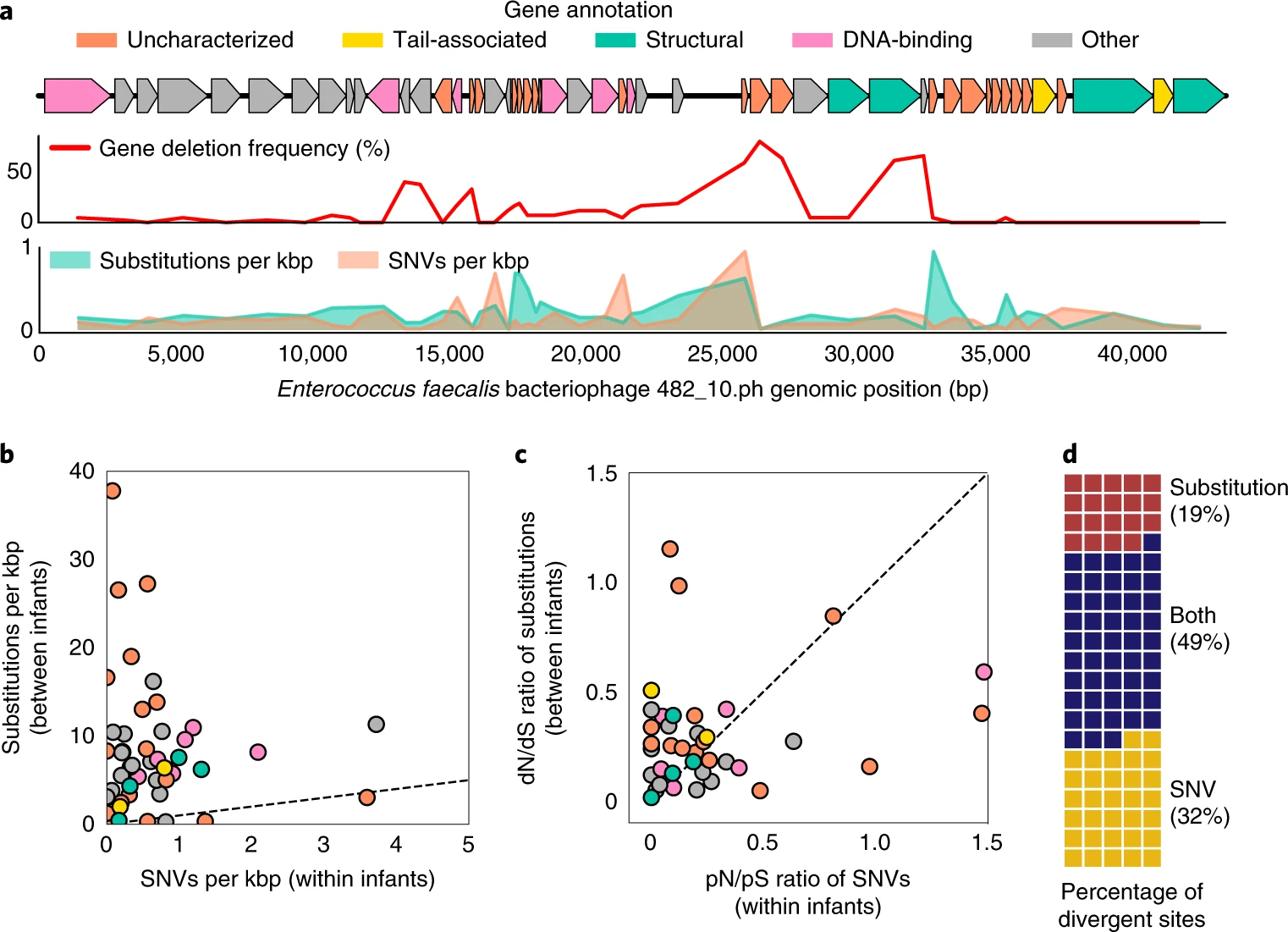
inStrain is python program for analysis of co-occurring genome populations from metagenomes that allows highly accurate genome comparisons, analysis of coverage, microdiversity, and linkage, and sensitive SNP detection with gene localization.
-
Source code is available on GitHub
-
Manual, installation instructions, and expected output are at available at ReadTheDocs
-
Publication is available in Nature Biotechnology and on bioRxiv
dRep
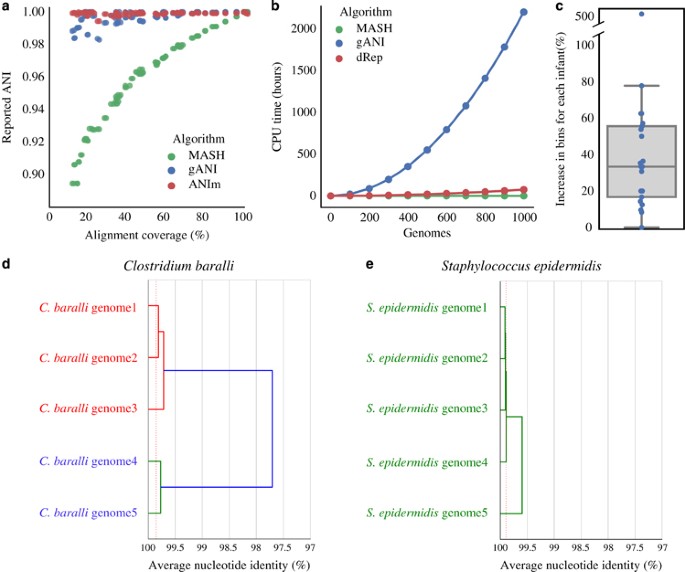
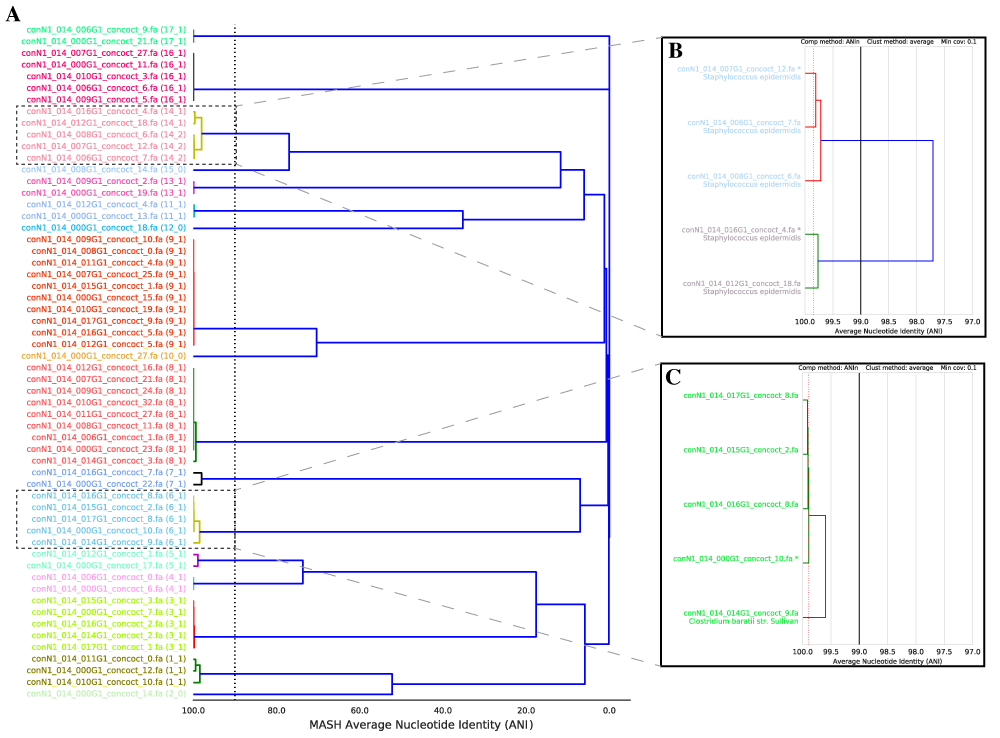
dRep is a python program for rapidly comparing large numbers of genomes. dRep can also “de-replicate” a genome set by identifying groups of highly similar genomes and choosing the best representative genome for each genome set.
-
Source code is available on GitHub
-
Manual, installation instructions, and API are at available at ReadTheDocs
-
Publication is available at The ISME Journal and on bioRxiv